Electrostatic fluctuation around acetylcholinesterase dimer
Stanislaw T. Wlodek (University of Houston) and J. Andrew McCammon
(University of California at San Diego)
Stanislaw T. Wlodek (University of Houston) and J. Andrew McCammon
(University of California at San Diego)

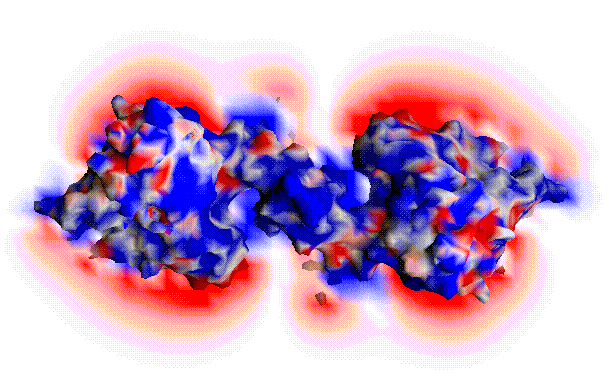
The correlation of electrostatic potentials between the crystal structures of the wild type and the ``neutral mutant'' of acetylcholinesterase (AChE). In the red region, the two types have opposite sign; in the blue region, the two types have the same sign.
Animation of electrostatic potential variations during an MD simulation of wild-type AChE (left); animation of electrostatic potential variations during an MD simulation of AChE ``neutral mutant'' (right).
The first animation shows the variations of the electrostatic potential around the enzyme acetylcholinesterase that arise from the thermal motion of the enzyme. The structural fluctuations of the enzyme are from a molecular dynamics simulation [Wlodek et al., 1997]. The electrostatic potential was determined for a series of snapshots from the simulation by solution of the Poisson-Boltzmann equation with the UHBD software [Madura et al., 1995]. The electrostatic calculations are for conditions of room temperature, and physiologic ionic strength of 150 mM. The electrostatic potential is in units of kcal/(mol e).
The calculations suggest that the regions of negative potential (indicated with red) that surround the entrances to the two active sites of the dimeric enzyme, are only marginally disturbed by protein motion. The regions of positive potential appear to be significantly more fluxional. The negative potential around the active sites of acetylcholinesterase is known to contribute very substantially to the high speed of the enzyme, by steering the positively-charged substrate acetylcholine to the active sites [Radic et al., 1997]. The relative stability of the regions of negative potential may therefore enhance the effectiveness of this electrostatic steering.
This work was supported in part by grants from NSF, NIH, and the San Diego Supercomputer Center.